Demo
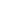
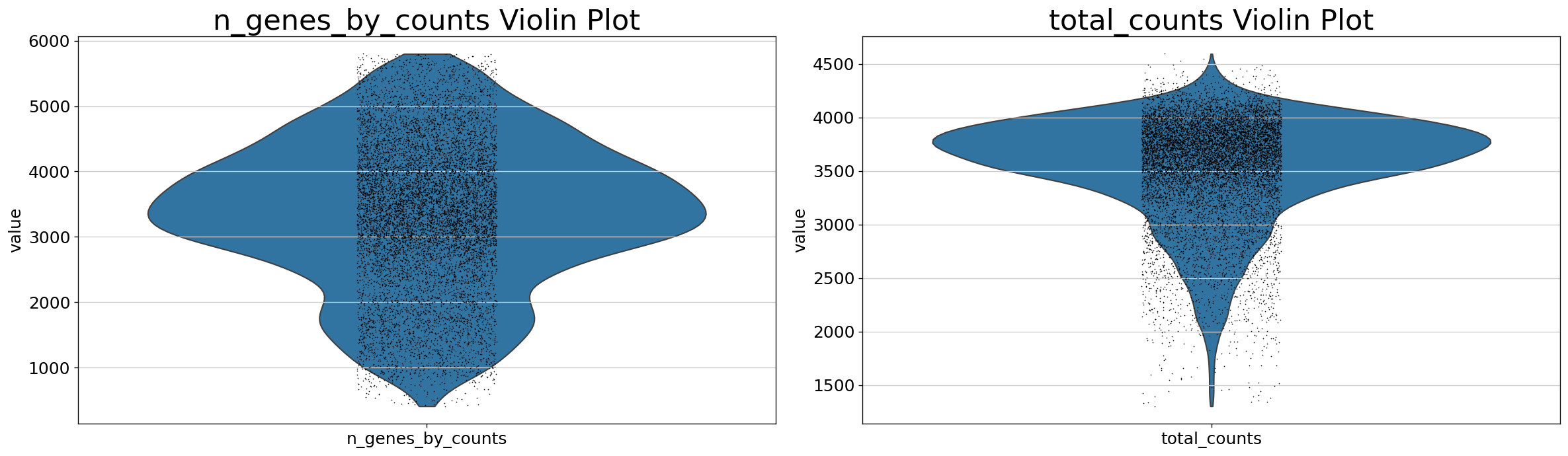
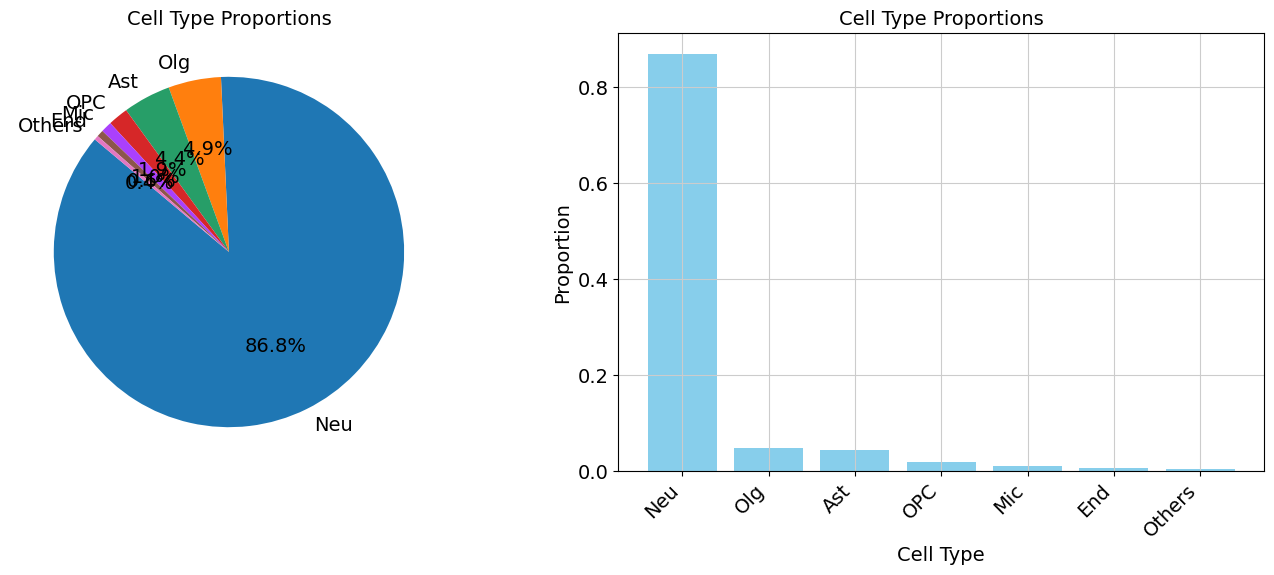
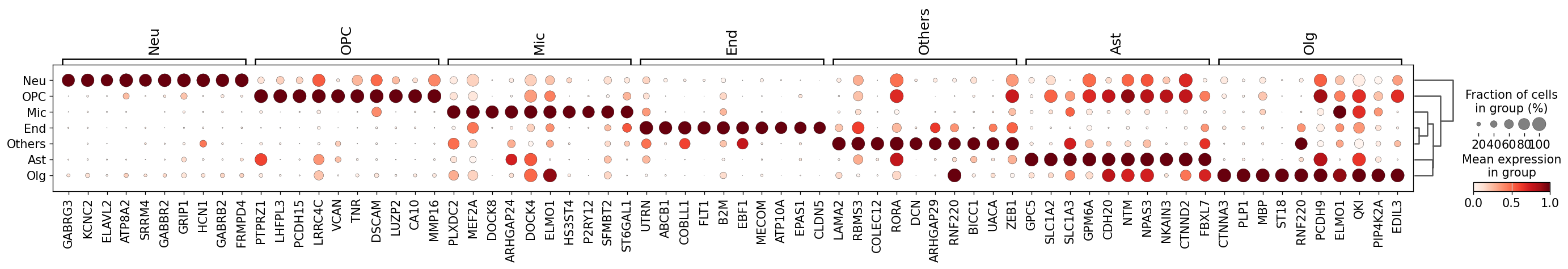
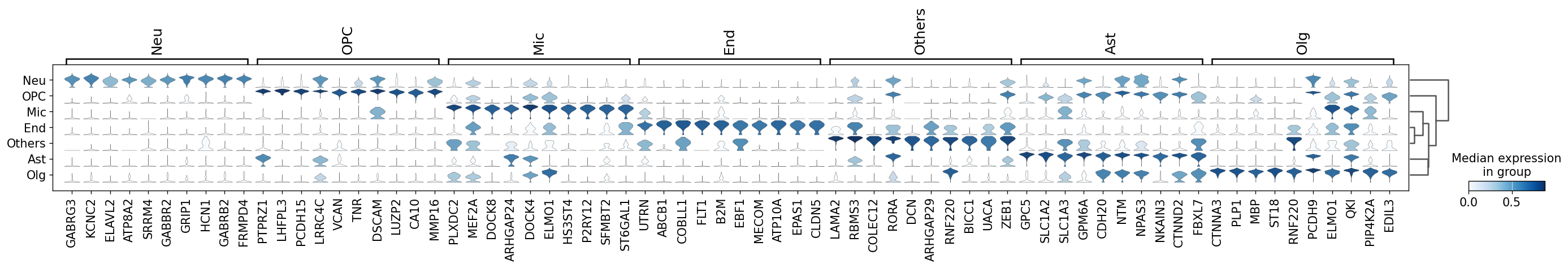
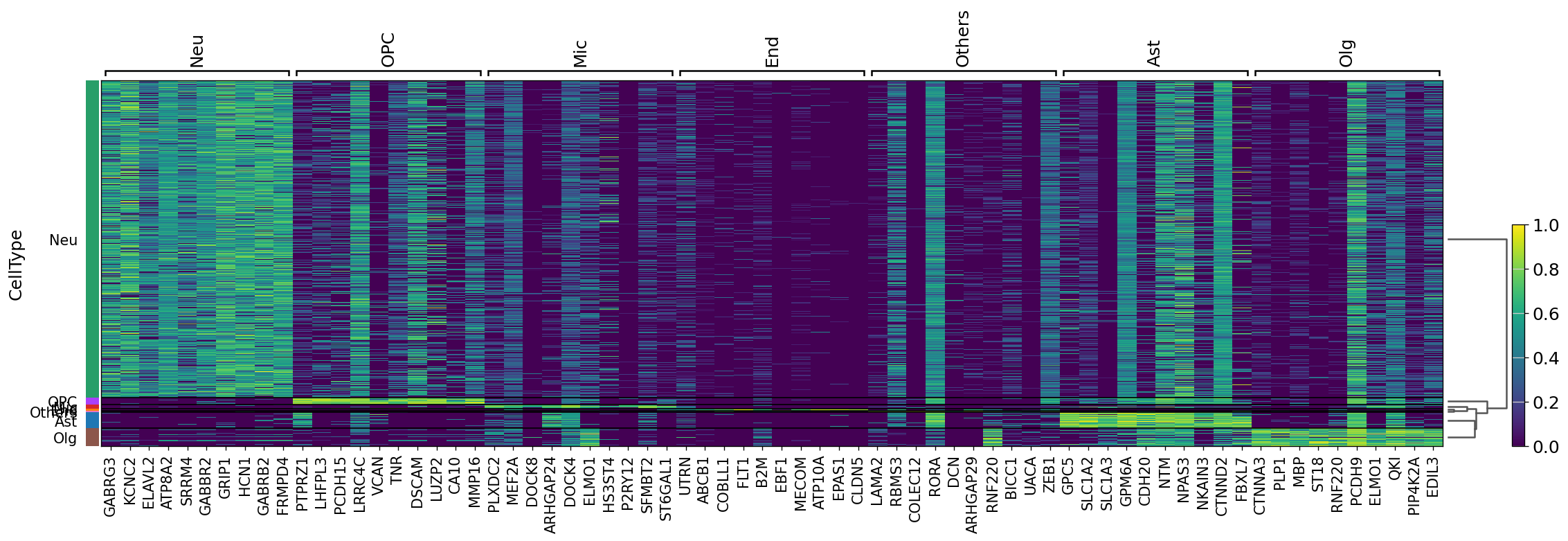
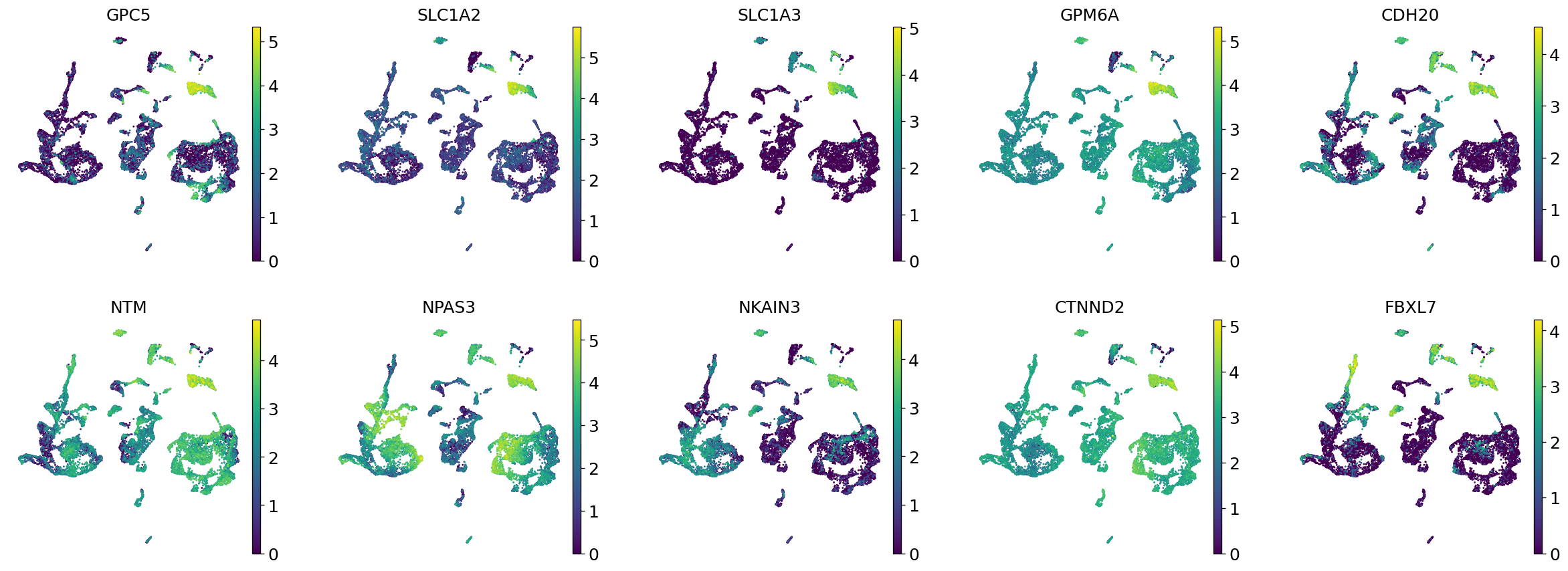
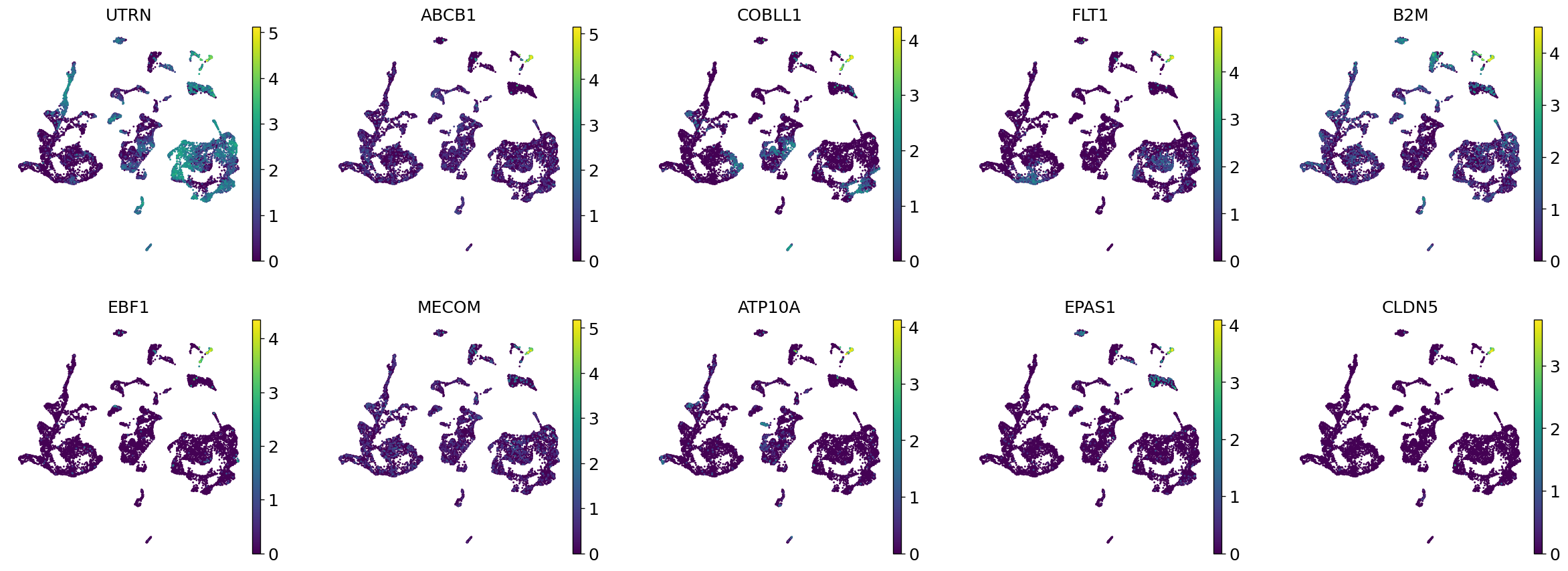
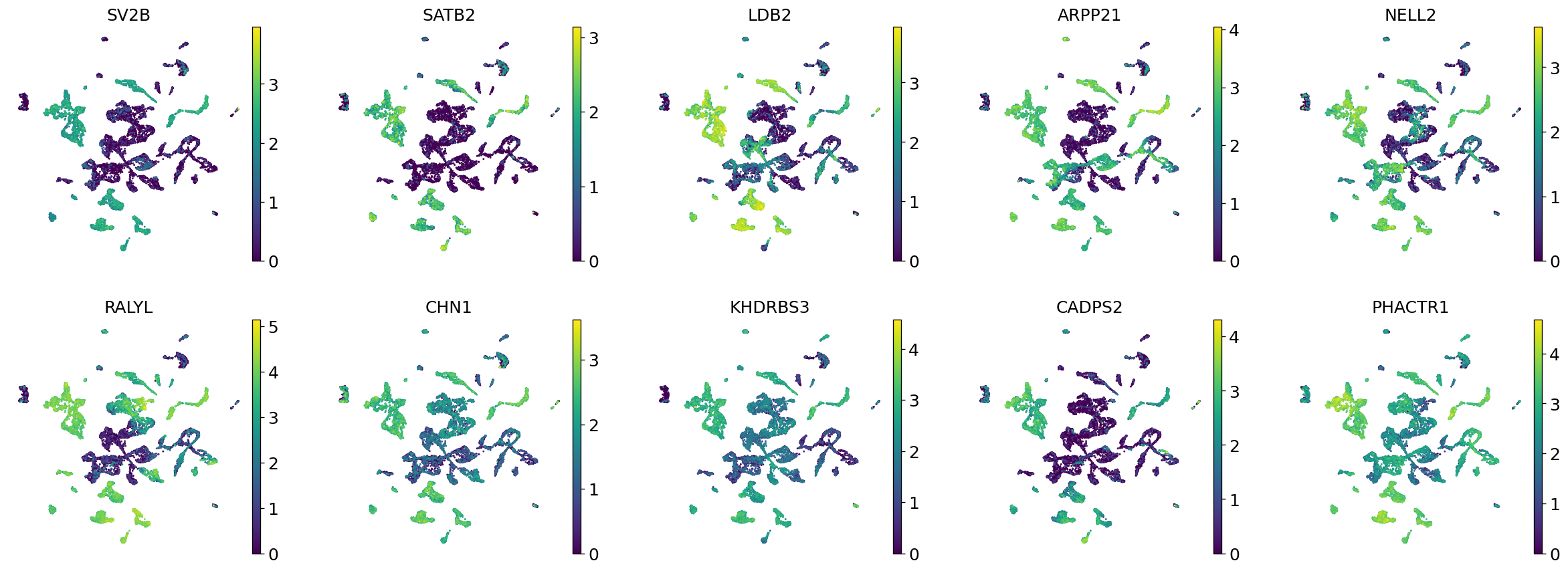
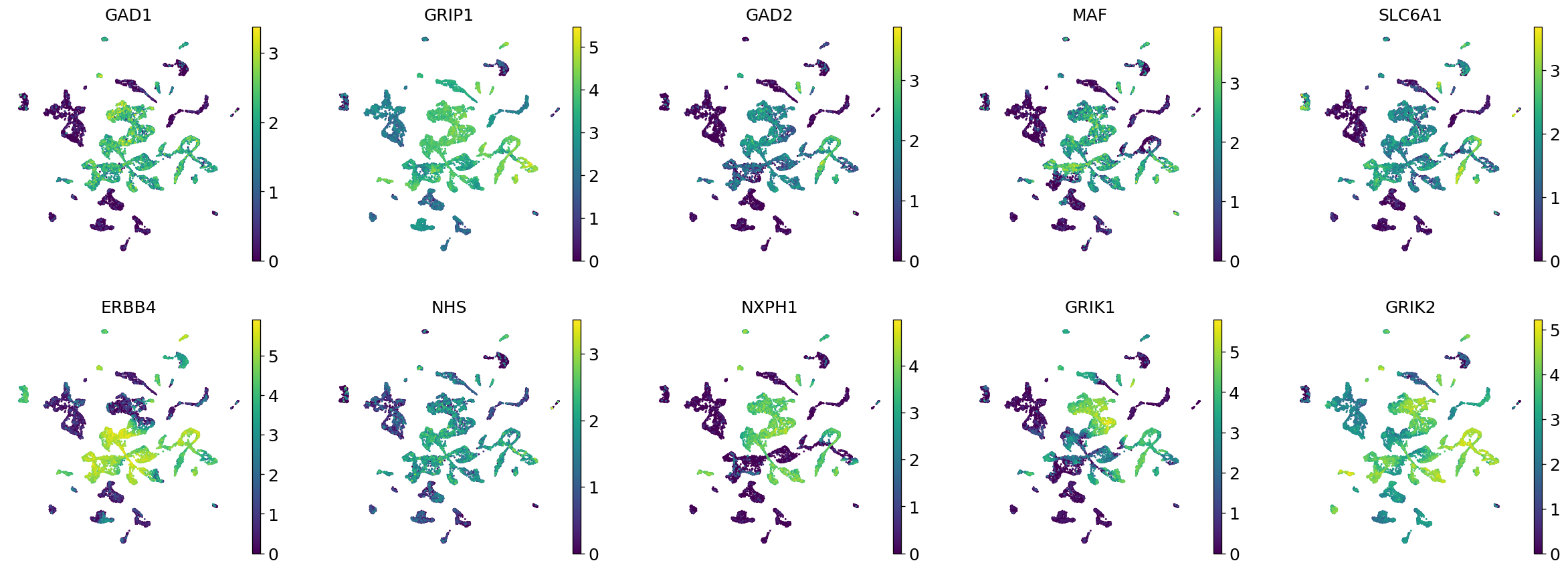
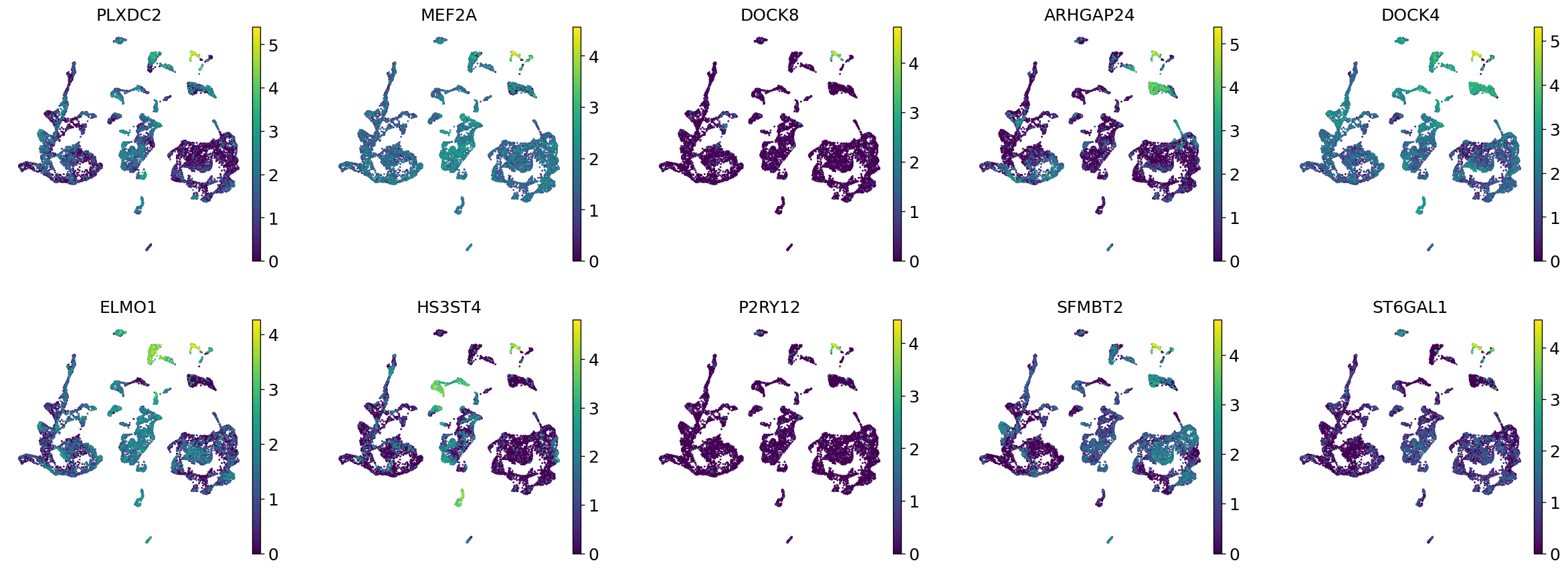
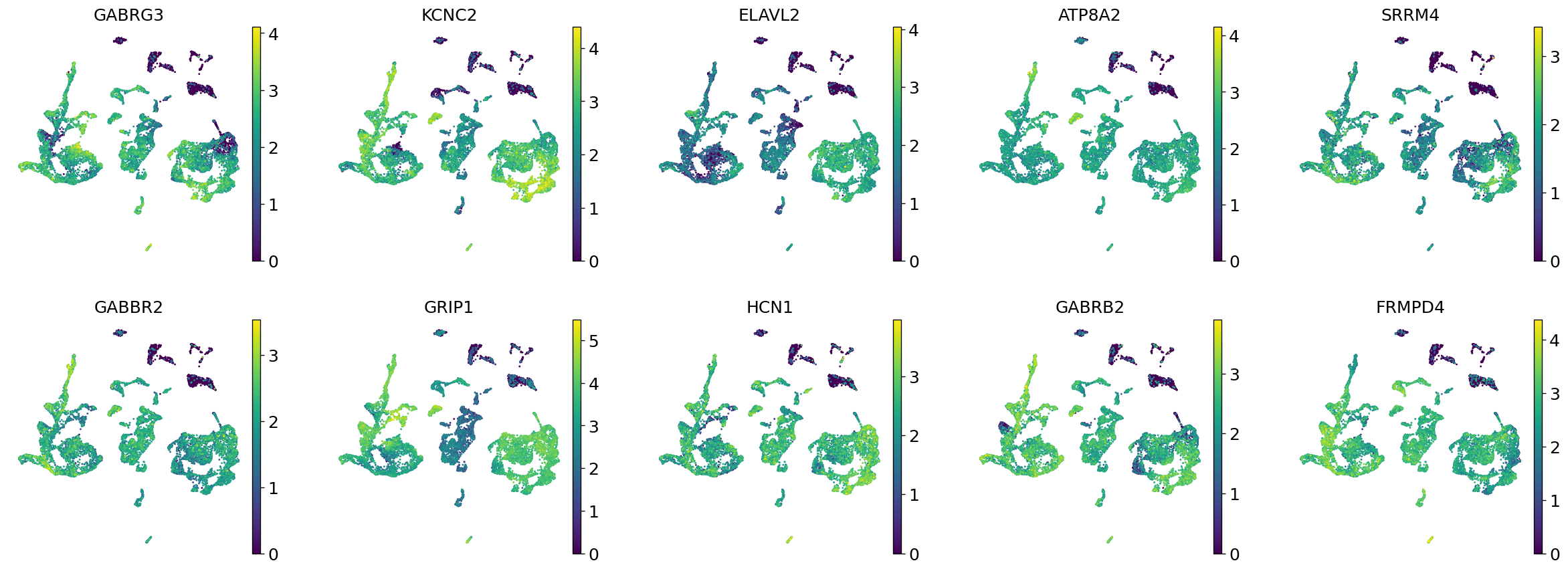
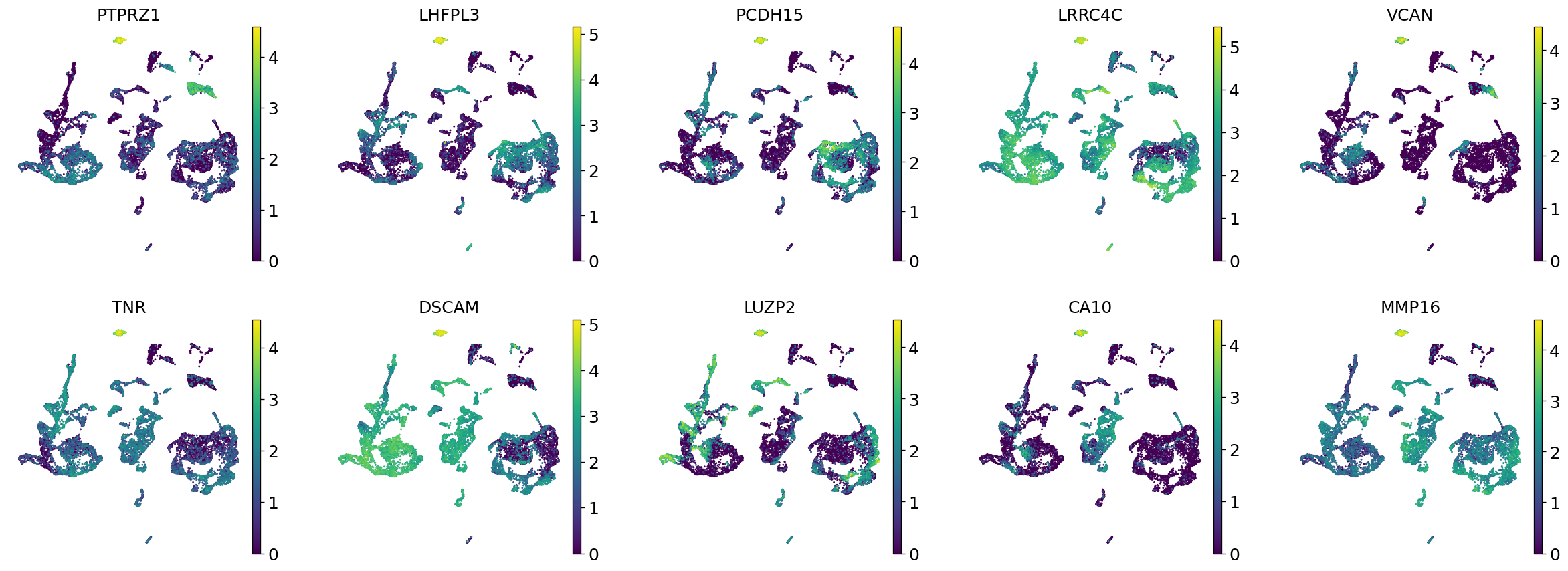
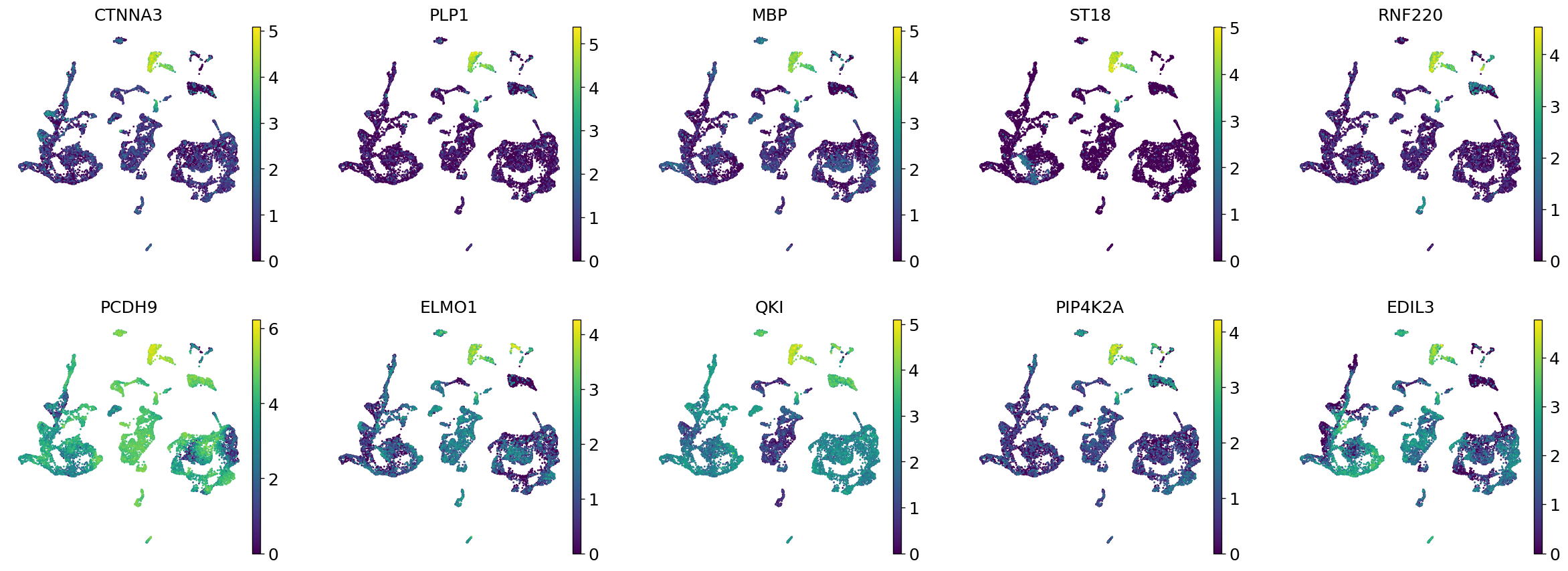
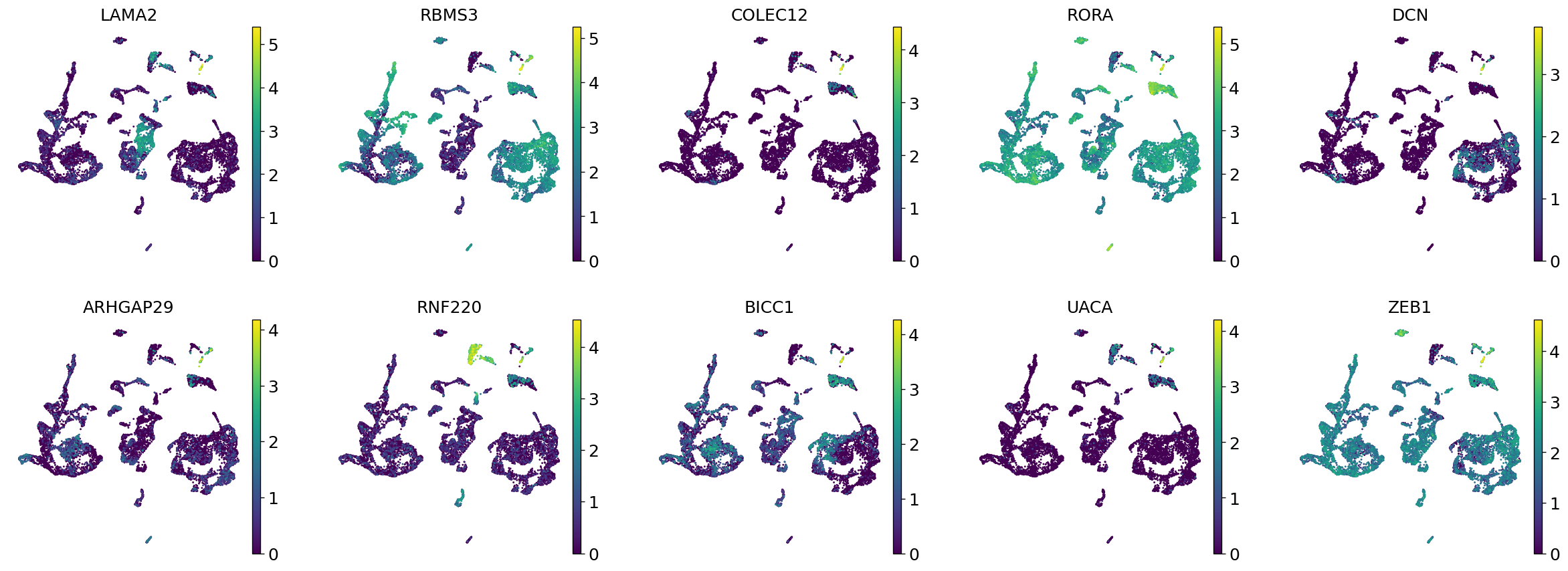
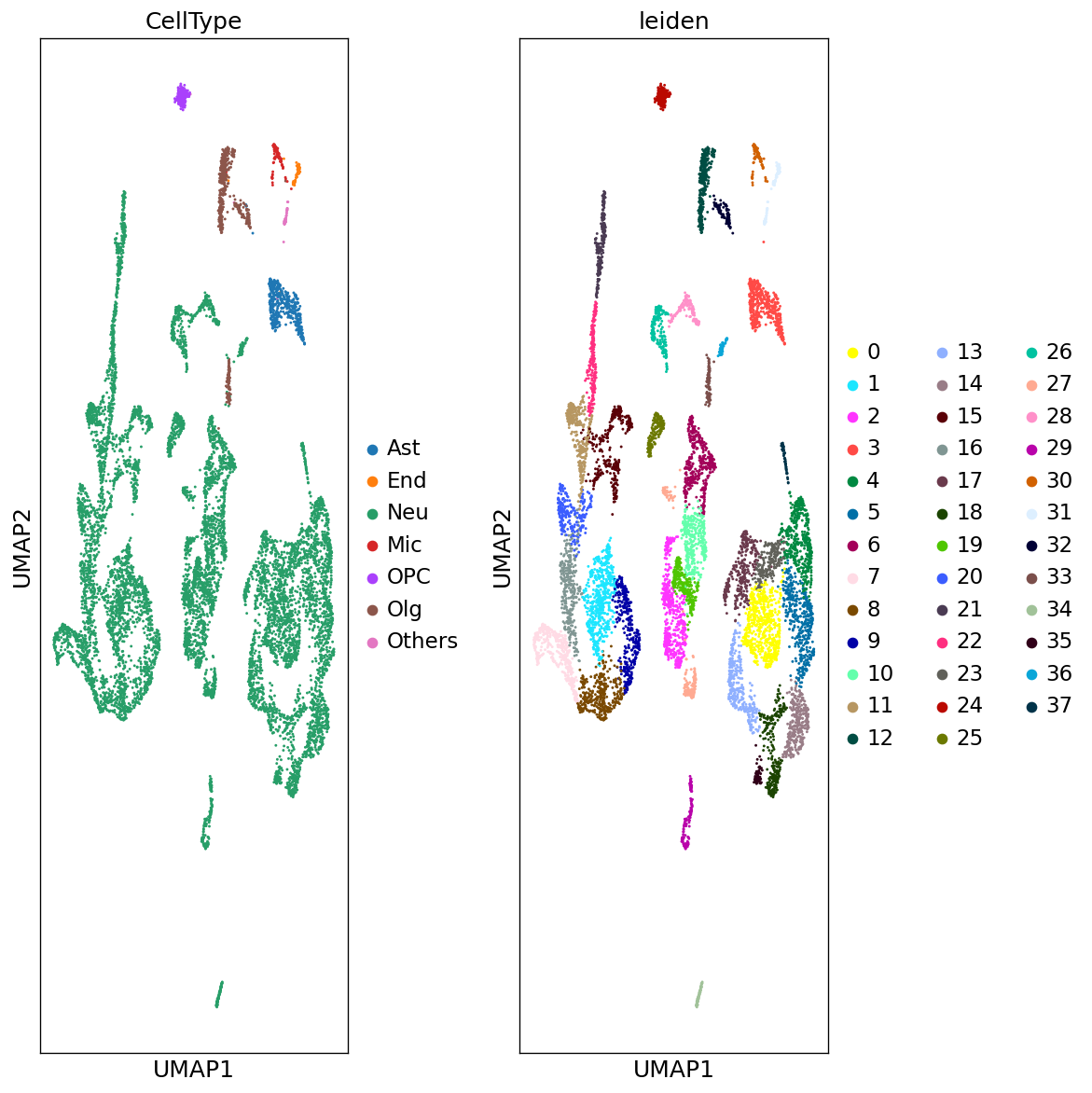
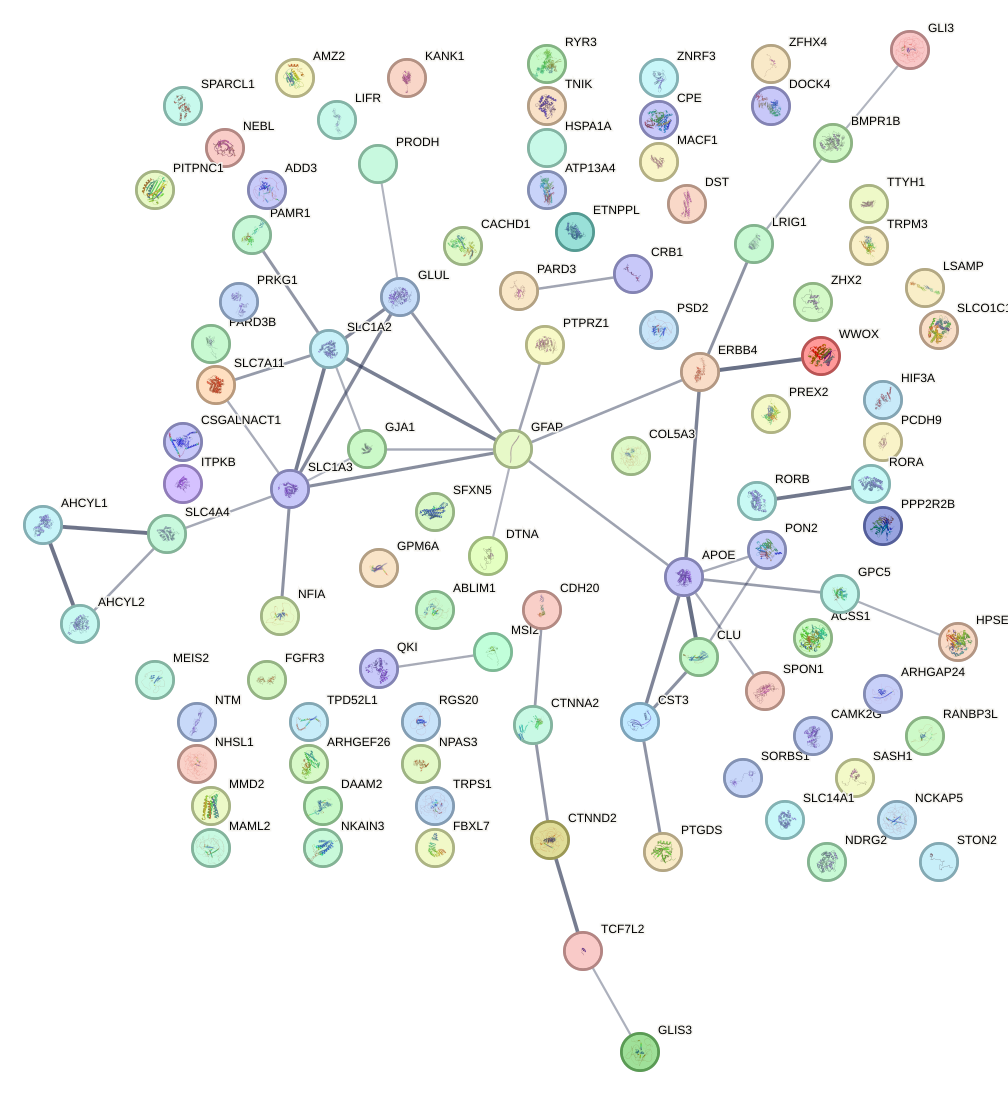
Demo serves as an interactive showcase of BrainScope’s data analysis capabilities, providing users with example analyses and visualizations to help them better understand how to utilize the platform effectively. This page features sample datasets and analytical workflows that highlight the core functionalities of BrainScope, including single-gene and gene-set expression analysis, cell-type marker exploration, and comparative transcriptomic studies across species, brain regions, and conditions. Users can navigate through different sections such as DataInfo & Gene: Explore dataset metadata, gene expression patterns, and cell marker expression at the single-gene level using Cytopedia; ShinyAtlas: Visualize interactive single-cell and spatial transcriptomic atlases; HomoGene: Compare homologous gene expression across different species; GeneSets & TF: Perform gene set and transcription factor enrichment analysis; Stats & Anno: Access statistical summaries and dataset annotations; TimeScale Review: Investigate evolutionary and developmental gene expression trends; Enrichment & Download: Conduct functional enrichment analysis and download processed results. By presenting step-by-step analytical demonstrations, the Demo page provides an intuitive entry point for new users while serving as a reference for advanced researchers looking to leverage BrainScope’s full analytical power.
VIEW HELP